|
All chromatograms were blasted to check for contamination using SpeciesTrilobodrilus windanseasp. The results confirmed that these transcripts are in fact differentially expressed between sporulated and unsporulated oocysts. The sequence reads were assembled into contigs using the computer pro- gram Sequencher for Macintosh (Gene Codes. Chromatogram reading and contig assembly were performed in Sequencher 4.10.1 (GeneCodes Corporation, Ann Arbor, MI, USA). The expression of 4 transcripts obtained from the subtracted cDNAs was confirmed by quantitative reverse transcriptase–polymerase chain reaction. Contig Organism: Pseudomonas aeruginosa MH38 (g-proteobacteria) Submitter: CEBITEC Date: 7 Assembly type: Assembly level: Contig Genome representation: full GenBank assembly accession: GCA000689435.1 (latest) RefSeq assembly accession: GCF000689435. Approximately half of the unique transcripts generated from sporulated oocysts are also expressed by sporozoites and merozoites, whereas the expression of most (79%) of the transcripts from unsporulated oocysts has not yet been detected at other stages of development. The file size of the latest downloadable installer is 300.6 MB. A total of 163 unique sequences were found, and the majority of these (64%) represent novel genes with no significant homology to the proteins in GenBank. Of these, 225 clones were isolated from cDNA of sporulated oocysts and 274 from unsporulated oocysts. 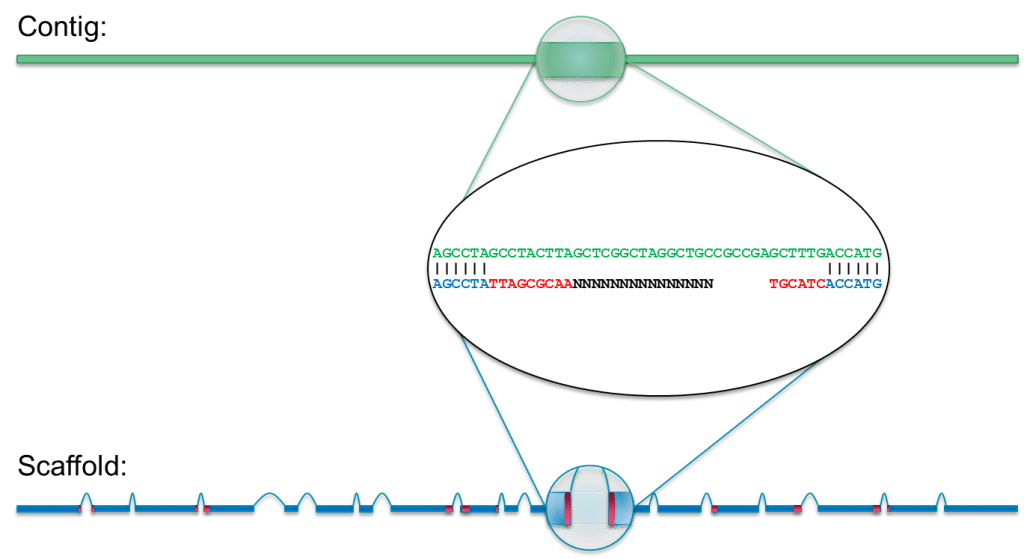
This allows you to compare a feature or a. To characterize the genes expressed by Eimeria tenella oocysts, the sequence of 499 expressed sequence tags (ESTs) was obtained from complementary DNA (cDNAs) enriched for transcripts expressed by unsporulated or sporulated oocysts. The Reference Sequence function sets the base numbering and the orientation of the contig you are about to assemble.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
